RODEO
RODEO (Rapid ORF Description & Evaluation Online) is an algorithm to help biosynthetic gene cluster (BGC) analysis, with an emphasis on ribosomal natural product (RiPP) discovery.
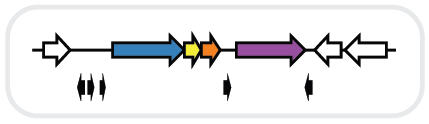
RODEO (Rapid ORF Description & Evaluation Online) is an algorithm to help biosynthetic gene cluster (BGC) analysis, with an emphasis on ribosomal natural product (RiPP) discovery.

RODEO evaluates one or many genes, characterizing a gene neighborhood based on the presence of profile hidden Markov models (pHMMs). Because RiPP precursor peptides are small and often evade annotation in sequence databases, RODEO can also manually translate small ORFs (open reading frames). A combination of support vector machine (SVM) learning and motif analysis can be used to scan unannotated intergenic regions for RiPP precursors.
With RODEO, thousands of gene clusters can be annotated in a single analysis. We have used RODEO to survey the biosynthetic landscape of certain RiPPs, annotate sequence-similarity networks and phylogenetic trees, decipher evolutionary relationships between enzymes & substrates, and track protein-protein co-occurrence. RODEO works on bacterial, archaeal, and fungal genomes.
To use RODEO to analyze a gene neighborhood, you will need to submit a NCBI protein accession (or list of accessions) to our web tool. By default, RODEO will annotate the gene neighborhood of your accession using the PFAM HMM database. When you submit the accession you have the choice to also have the gene neighborhood annotated using TIGRFAM, RREFam or uploaded custom HMMs. You may also choose to analyze the gene neighborhood for certain RiPP classes, which will search the gene neighborhood for precursor peptides from that RiPP class. If you have proprietary or unpublished genomes you would like to analyze by RODEO,please contact the Mitchell lab to discuss potential collaberations.
If you have used RODEO, please consider citing the publication relevant to your use:
Initial publication: Tietz, J.I.; Schwalen, C.J.; Patel, P.S.; Maxson, T.; Blair, P.M.; Tai, H.C.; Zakai, U.Z.; Mitchell, D.A. "A new genome mining tool redefines the lasso peptide biosynthetic landscape." Nat. Chem. Biol., 13: 470-478 (2017). doi:10.1038/nchembio.2319
Lasso peptide Scoring: Tietz, J.I.; Schwalen, C.J.; Patel, P.S.; Maxson, T.; Blair, P.M.; Tai, H.C.; Zakai, U.Z.; Mitchell, D.A. "A new genome mining tool redefines the lasso peptide biosynthetic landscape." Nat. Chem. Biol., 13: 470-478 (2017). doi:10.1038/nchembio.2319
Thiopeptide/Pyritide Scoring: Schwalen, C.J.; Hudson, G.A.; Kille, B.; Mitchell, D.A. "Bioinformatic expansion and discovery of thiopeptide antibiotics." J. Am. Chem. Soc., 140: 9494-9501 (2018). doi:10.1021/jacs.8b03896
Sactipeptide/Ranthipeptide Scoring: Hudson, G.A.; Burkhart, B.J.; DiCaprio, A.J.; Schwalen, C.; Kille, B.; Pogorelov, T.V.; Mitchell, D.A. "Bioinformatic mapping of radical S-adenosylmethionine-dependent ribosomally synthesized and post-translationally modified peptides identifies new Cα, Cβ, and Cγ-linked thioether-containing peptides." J. Am. Chem. Soc., 141: 8228-8238 (2019). doi:10.1021/jacs.9b01519
Lanthipeptide Scoring: Walker, M.C.; Eslami, S.M.; Hetrick, K.J.; Ackenhusen, S.E.; Mitchell, D.A.; van der Donk, W.A. "Precursor peptide-targeted mining of more than one hundred thousand genomes expands the lanthipeptide natural product family." BMC Genomics, 21, 387-403 (2020). doi.org/10.1186/s12864-020-06785-7
Linaridin Scoring: Georgiou, M.A.; Dommaraju, S.R.; Guo, X.R.; Mast, D.H.; Mitchell, D.A. "Bioinformatic and reactivity-based discovery of linaridins." ACS Chem. Biol., 15: 2976-2985 (2020). doi.org/10.1021/acschembio.0c00620
Graspetide Scoring: Ramesh, S.; Guo, X.; DiCaprio, A.J.; De Lio, A.M.; Harris, L.A.; Kille, B.L.; Pogorelov, T.V.; Mitchell, D.A. "Bioinformatics-guided expansion and discovery of graspetides." ACS Chem. Biol., 16: 2787-2797 (2021). doi.org/10.1021/acschembio.1c00672
RREFams: Kloosterman, A. M.; Shelton, K. E.; van Wezel, G. P.; Medema, M. H.; Mitchell, D. A. RRE-Finder: A Genome-Mining Tool for Class-Independent RiPP Discovery. mSystems 2020, 5 (5), e00267– 20, DOI: 10.1128/mSystems.00267-20